|
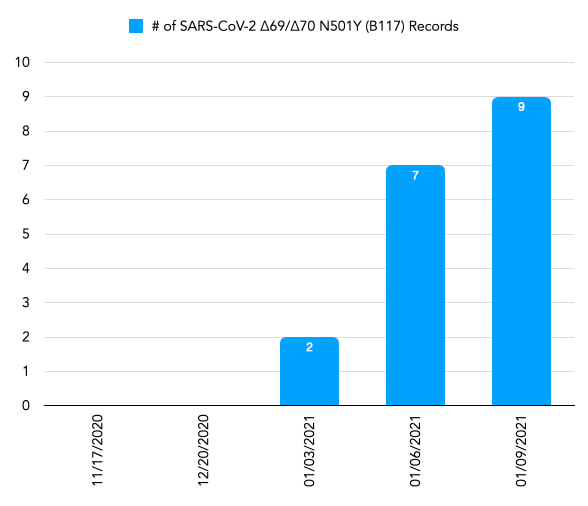
Over the past several of weeks news reports have come out describing the appearance of two variants of SARS-CoV-2 that are thought to be more transmissible compared to the variants that are currently circulating in the community. One of these variants, known as 20B/501Y.V1 VOC 202012/01 or B.1.1.7) was first identified in the United Kingdom in September of 2020. SARS-Cov-2-B117 has multiple mutations including a deletion of amino acids 69 and 70 and the substitution of Y for N at position 501 (N501Y) in the spike protein. The second variant was identified in South Africa in October of 2020 and is known as 20C/501Y.V2 or B1.351). SARS-CoV-2-B1351 also contains multiple mutations in the spike protein including the N501Y substitution, but unlike SARS-CoV-2-B117 does not contains the 69/70 deletion. I wanted to extend the analysis of SARS-CoV-2 spike protein mutations we started in the CHEM 440 class to identify SARS-CoV-2-B117 variants in the NCBI database. During our first pass through the spike protein sequences in the NCBI database we limited our analysis to sequences that were the same length as the reference spike protein sequence. This made it easy to identify amino acid mutations. Since SARS-CoV-2-B117 has two amino acid deletions in the spike protein, and is therefore shorter than the reference protein sequence, it is not possible to find mutations using the python code we had written. To analyze sequences that are shorter than the reference sequence, I updated the code to carry out pairwise alignments of each database entry with the reference sequence using the Biopython pairwise2 module. Each alignment was checked to see if the query (i.e. the database entry spike protein) sequence contained a deletion at positions 69 and 70 and a N to Y substitution at position 501. Database entries with these characteristics were identified as samples that contained the SARS-CoV-2-B117 variant. Analysis of the SARS-CoV-2 spike protein sequences in the NCBI database resulted in the following findings: The NCBI database downloaded on 11/17/2020 nor the one downloaded on 12/20/2020 contained any entries that were identified as carrying the SARS-CoV-2-B117 variant. Two samples were identified as carrying the SARS-CoV-2-B117 variant on 1/03/2021. Of these two samples one was collected in Colorado on 12_24_2020 and deposited in the NCBI database on 12_30_2020 while the other was collected in San Diego, CA on 12_29_2020 and deposited in the NCBI database on 12_30_2020. By 1_6_2021 an additional five samples containing the SARS-CoV-2-B117 variant was deposited into the database. Of these sample one was collected in Florida, three were collected in California and one was collected in Saratoga County, New York. All five samples were collected between 12_19_2020 and 12_24_2020 and deposited in to the NCBI database on 1_1_2021. Two additional samples, collected in Italy, containing the B117 variant was deposited in to the database after 1_6_2021.
Given the nature of sample collection and sequencing it is highly unlikely that the number of samples identified here is any indication of the community prevalence of the B117 variant. As public health officials increase testing and sequencing efforts the number of samples containing the B117 variant is expected to increase rapidly. I will keep analyzing the NCBI database every three to four days to see if we can capture this increase. Interestingly in an article published in April of 2020 (Wan Y, Shang J, Graham R, Baric RS, Li F. 2020. Receptor recognition by the novel coronavirus from Wuhan: an analysis based on decade-long structural studies of SARS coronavirus. J Virol 94:e00127-20. https://doi.org/10.1128/JVI.00127-20.) Wan et.al. suggested that mutations at residue 501 of the spike protein could "significantly enhance the binding affinity between 2019-nCoV RBD and human ACE2. Thus, 2019-nCoV evolution in patients should be closely monitored for the emergence of novel mutations at the 501 position (to a lesser extent, also the 494 position)". Early in the pandemic SARS-CoV-2 was known as 2019-nCoV.
0 Comments
Leave a Reply. |
ArchivesCategoriesAuthorHi, my name is Sajith Jayasinghe and I am faculty member in the department of Chemistry and Biochemistry at California State University, San Marcos, CA. My faculty information can be found at https://faculty.csusm.edu/jayasinghe/index.html and details of my research efforts can be found at public.csusm.edu/jayasinghe/ |
Site powered by Weebly. Managed by Bluehost
 RSS Feed
RSS Feed